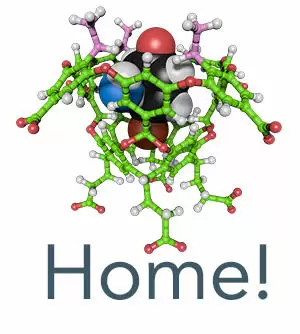
Yao, W.; Wang, K.; Wu, A.; Reed, W. F.; Gibb, B. C. Anion binding to ubiquitin and its relevance to the Hofmeister effects. Chemistry Science 2021, 12, 320-330.
Although the non-covalent interactions between proteins and salts contributing to the Hofmeister effects have been generally mapped, there are many questions regarding the specifics of these interactions. We report here studies involving the small protein ubiquitin and salts of polarizable anions. These studies reveal a complex interplay between the reverse Hofmeister effect at low pH, the salting-in Hofmeister effect at higher pH, and six anion binding sites in ubiquitin at the root of these phenomena. These sites are all located at protuberances of preorganized secondary structure, and although stronger at low pH, are still apparent when ubiquitin possesses no net charge. These results demonstrate the traceability of these Hofmeister phenomena and suggest new strategies for understanding the supramolecular properties of proteins.